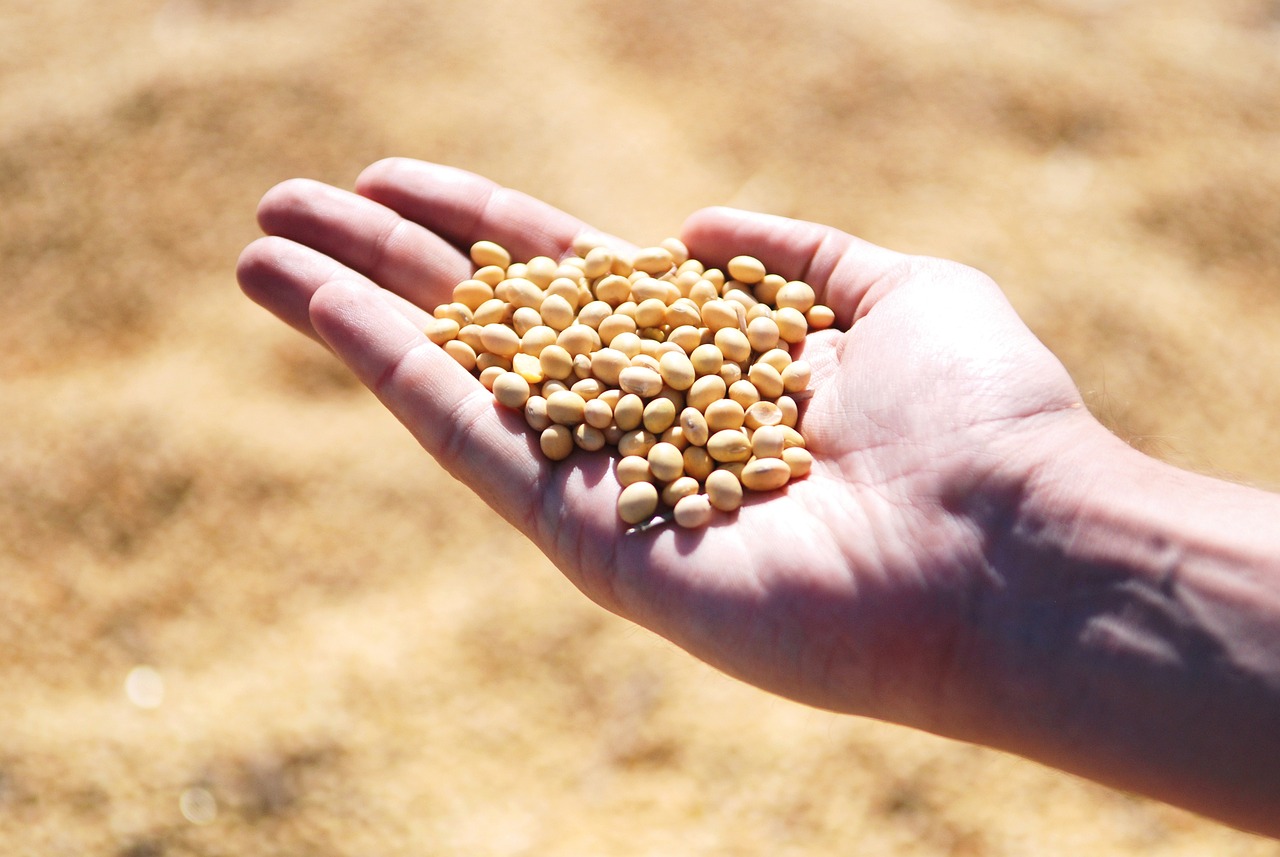
: The project will empower soybean researchers and breeders to directly select for otherwise hidden genetic variation in genes that can be targeted for variety development.
Soybean is an important crop, both in terms of its protein production and its use as a biodiesel. A new study led by the University of Illinois Urbana-Champaign and the U.S. Department of Energy Joint Genome Institute (JGI) plan to sequence 400 soybean genomes to develop a “pangenome”: a comprehensive characterization of all the useful diversity in the genome. The project aims to contribute knowledge that ultimately helps build more robust and resilient soybean crop that performs to maximum potential.
The project aims to drastically improve the current soybean reference genome, according to a release. Currently the reference genome does not reflect the real diversity of the soybean crop. This limitation is not unlike the first human genome, which was made up of only Caucasian individuals.
“There’s an increasing effort to have the reference human genome reflect all of the variation in people. We think there are equally big reasons to do the same thing in crops,” Hudson said in the release. “But it’s hard to locate the missing diversity by any other means than sequencing more genomes.”
The soybean pangenome project will sequence and analyze at least 50 soybean genomes from cultivated lines and wild relatives at reference quality, the gold standard of modern sequencing. The project will also sequence an additional 350 genomes as high-quality drafts by the JGI. Included in the sequencing will be perennial relatives and lines selected to yield in harsh conditions. Focusing on these kinds of diversity will position the agriculture industry as more climate-resilient. The team intends to solicit help from the soybean breeding community, from academics to industry partners, to determine which lines should be included in the project.
“There have been soybean pangenome efforts before, but this will be a big step forward. We want to identify all of the variation present within this diverse set of cultivated soybeans. Knowing details of all of the genetic variation should very much enhance and speed up the ability of crop breeders and biotechnology experts to identify important genes and incorporate them into better crops,” said project leader Matt Hudson in the release, professor in the Department of Crop Sciences, part of the College of Agricultural, Consumer and Environmental Sciences (ACES) at U. of I. Hudson is uniquely positioned to add value to this project, as he also serves as co-director of the Center for Digital Agriculture, science integration chair for the Center for Advanced Bioenergy and Bioproducts Innovation (CABBI), and faculty affiliate at the Carl R. Woese Institute for Genomic Biology.
The project depends on the unique skills and infrastructure of both partners. Hudson and his team will begin the process by selecting and growing soybean lines. They will then ship extracted DNA to JGI for long-read sequencing. Once sequenced, Hudson’s team and partners at U. of I.’s AIFARMS will lead analyzing the output.
The project will ultimately “enable deep analysis of the evolution and domestication of modern soybean and empower soybean researchers and breeders to directly select for otherwise hidden genetic variation in genes that can be targeted for variety development. As soybean is becoming increasingly important as a worldwide crop, as well as being a key bioenergy crop, this project will have global impact and be particularly relevant to U.S. agriculture,” said Hudson in the release.