The first soybean cyst nematode pangenome reveals how genetically diverse the pest is, offering new insight into how it overcomes resistance and how researchers might stay ahead.
Few pests eat away at farm profitability like Soybean cyst nematode (SCN). Responsible for at least $1.5 billion in yield losses each year, it remains soybean’s most damaging pest in the United States.
For decades, the industry’s most reliable defense has been genetic resistance. But that protection is starting to weaken as SCN populations evolve.
Now a multi-institution research team led by University of Illinois Urbana-Champaign has produced the first pangenome for the pest — a comprehensive genetic map that reveals just how diverse and adaptable SCN populations really are. The work was supported by the North Central Soybean Research Program and the Illinois Soybean Association.
“The SCN pangenome allows us to do population genetic analysis of nematodes to understand how they’re overcoming resistance,” says Matt Hudson, professor in the Department of Crop Sciences at Illinois and senior author of the study, in the university news release.
Hudson says the pangenome could eventually support tools that help farmers identify which soybean resistance sources are most effective in a given region. In the longer term, it may also help researchers design strategies to suppress the most virulent SCN populations before they cause yield losses.
What a Pangenome Reveals
A pangenome captures the full genetic diversity of a species rather than relying on a single reference genome.
“It’s a modern approach scientists use to catalog the genetic diversity of a species. For decades, the standard practice was to choose a single individual and treat it as the reference. Imagine choosing a single person to represent all of the genetic diversity of the human species; that’d be impossible,” says Lucas Borges dos Santos, a doctoral student in the Informatics Program at Illinois.
Hudson and his colleagues have previously led a soybean pangenome project analyzing more than 400 plants, resulting in multiple studies that describe the crop’s genetic diversity. Building a pangenome for SCN proved significantly more difficult.
Cracking a Difficult Genetic Puzzle
Collecting high-quality DNA from SCN presents unique challenges.
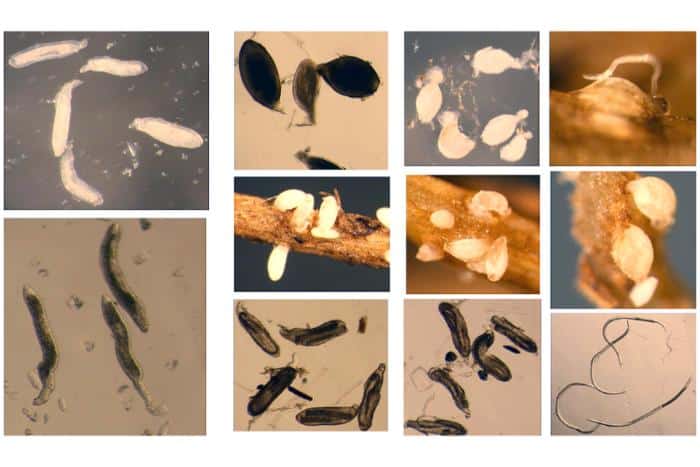
“One problem with SCN is that they’re either living in soybean roots, swimming around in the soil, or they’re eggs. It’s hard to extract DNA from adults, so we typically try eggs,” Hudson says. “But with their hard exterior, they’re like little cans of beans in the soil. That lets them survive for decades, but makes it really hard to extract DNA.”
Earlier approaches required scientists to freeze-dry SCN eggs and crush them into powder. The mechanical force shattered the DNA into fragments too small for high-quality genome assembly.
Several years ago, study co-author Kim Walden of the Roy J. Carver Biotechnology Center developed an enzyme mixture that gently dissolves the eggshell while preserving intact DNA. That breakthrough made the SCN pangenome possible. Walden passed away in 2024.
Mapping SCN Diversity
Researchers typically measure SCN virulence using the HG Type test, which infects seven soybean varieties and measures how successfully the nematodes reproduce. If SCN populations develop females on soybean lines 2, 5 and 7, for example, the population is classified as HG Type 2.5.7.
To build the pangenome, Borges dos Santos assembled genomes from nine SCN populations. Each population included several thousand individual nematodes and represented the full range of HG Types.
The results revealed extraordinary genetic variation within the pest.
“If you looked at the differences in DNA between humans and chimps, some of these worms would be humans and some of them would be chimps,” Hudson says. “The HG Types are very different from each other, and a lot of the genes are specific to each population. They’re very, very diverse.”
Why Resistance Breaks Down
That diversity helps explain why resistance can eventually fail. When SCN populations encounter resistant soybean varieties, early generations may struggle. Over time, however, the nematodes can draw on their extensive genetic diversity to activate virulence genes that allow them to reproduce again.
The pangenome provides researchers with a clearer picture of those genes and proteins, which could eventually support new strategies to control the pest.
“We’re looking at ways to alter the nematode populations themselves,” Hudson says. “For malaria mosquitoes, scientists can engineer and release male-sterile mosquitoes that reduce reproduction and spread of the disease. We have an Illinois Soybean Association grant along similar lines for SCN.”
A Foundation for Future Solutions
Although the research will not immediately change how farmers manage SCN, soybean leaders say the discovery provides an important new foundation for future solutions.
“Soybean cyst nematode remains the most damaging soybean pest in the U.S., costing farmers more than a billion dollars every year. This new pangenome finally gives researchers and breeders a full picture of SCN’s genetic diversity and adaptability that can deliver the deeper understanding we need to stay ahead of this pest,” says Stephanie Porter, outreach agronomist at the Illinois Soybean Association. “This research strengthens the foundation for developing more durable resistance and aligns directly with ISA’s commitment to protecting Illinois soybean yields.”
The study, Pangenome analysis of nine Soybean Cyst Nematode genomes reveals hidden variation contributing to diversity and adaptation, is published in BMC Genomics.