An international consortium of more than 80 researchers from over 30 institutions across nine countries is calling for a coordinated, worldwide effort to map gene expression across every cell type and developmental stage of wheat.
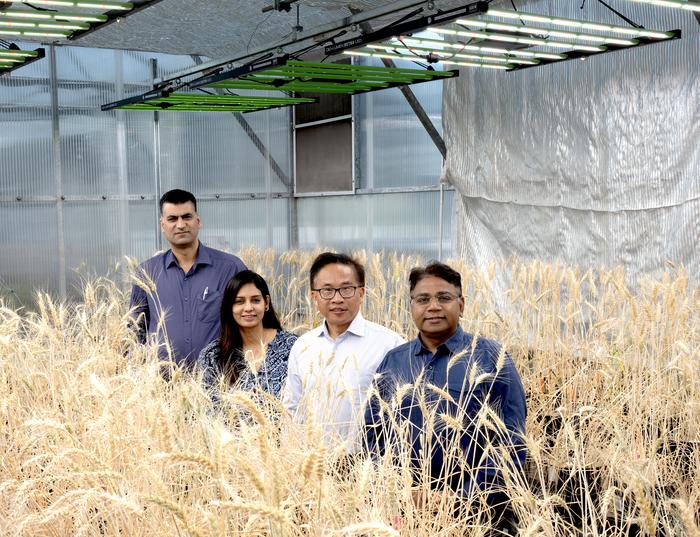
In a major perspective article published in Nature Genetics, the Wheat Spatial Omics Consortium (WSOC), led by researchers from Murdoch University’s Centre for Crop and Food Innovation (CCFI), Adelaide University, BGI Research and Xianghu Laboratory, present a strategic framework outlining how spatial omics technologies can transform wheat research by revealing how individual cell types respond to stress, regulate grain development and control yield-related traits.
Wheat provides roughly 35% of global dietary calories and feeds more than a third of the world’s population. With the global population projected to reach 9.8 billion by 2050, wheat production must increase by an estimated 60% to keep pace with demand; a target increasingly threatened by climate change, drought, heat stress and emerging diseases, according to a press release.
The paper details how spatial omics technologies, which map gene activity, proteins, and metabolites at the cellular level while preserving tissue architecture, can amplify the precision of genomics research to levels previously impossible with traditional bulk-tissue approaches.
The consortium has proposed a systematic experimental framework that goes well beyond current spatial transcriptomics research on wheat, calling for future efforts to encompass diverse wheat accessions, growth stages, tissues, stress treatments, and time points.
The authors also call for a multi-omics approach, which they consider essential for moving beyond description to a mechanistic understanding of how genes, proteins, and metabolic pathways interact within intact tissues. They add that artificial intelligence, including deep learning models for cell segmentation, graph-based spatial analysis and large language model-driven automation of research workflows, will be a key enabler for analysing the massive, multilayered datasets generated from this work, thereby dramatically narrowing the list of candidate genes and accelerating the discovery of elite genetic variants.
“Spatial omics will be vital for filling key knowledge gaps by identifying specific cells in stressed wheat tissues that express genes conferring tolerance,” Co-corresponding author and CCFI Director, Professor Rajeev Varshney, said. “With these insights, we can develop more targeted breeding strategies that accelerate the discovery of elite genetic variants, ensuring that our goal of feeding a growing global population is achievable.”