Watermelon is the quintessential summertime fruit, calling to mind warm afternoons, summer barbecues, and gatherings with friends and family. Its striped green rind and bright pink flesh are instantly recognizable; so are the crisp bite and rush of sweet juice that make each slice so familiar. What is less obvious is the biology behind those traits: the genetics shaping fruit color, texture, and sweetness through millions of years of natural selection and, more recently, deliberate breeding.
The watermelons commonly found in grocery stores represent only a small fraction of the fruit’s diversity. Each variety has a distinct set of physical traits, or phenotypes, shaped by differences in its genetic makeup, or genotype. Over time, breeding for preferred traits has narrowed the genetic diversity of cultivated watermelon, limiting its ability to withstand environmental stress and disease and raising concerns about its long-term sustainability. Wild watermelons, however, still preserve many valuable traits and may offer important tools for improving modern cultivars.
To better understand how genetic differences drive visible variation in watermelon, an international team led by Dr. Zhangjun Fei of the Boyce Thompson Institute developed a comprehensive watermelon super-pangenome, now published in Nature Genetics. This resource brings together 138 genomes from both wild and cultivated watermelons, revealing evolutionary relationships among individual genomes in unprecedented detail, according to a press release.
Traditional reference genomes are usually based on a single representative of a species. Pangenomes expand on that approach by incorporating multiple accessions within a species, but even they do not fully capture the diversity found across related species.
“We call ours a super-pangenome because we have incorporated accessions not just from one species, but from all seven extant species in the watermelon genus,” Fei explains. “The wild relatives provide abundant information for marker development and genomic prediction.”
Super-pangenomes are powerful tools for targeted crop breeding, helping researchers develop varieties that are better suited to society’s needs.
Watermelon genomes contain hundreds of megabases of DNA, and differences between individual genomes can range from small single nucleotide polymorphisms (SNPs) to large structural variants (SVs) spanning hundreds of thousands of bases. Because this watermelon super-pangenome is graph-based, it is especially effective at detecting SVs, which can have major effects on phenotype. By integrating these layers of information, Fei’s team created a detailed map of watermelon genetic diversity.
Using this resource, the team set out to connect specific genomic regions with important agricultural traits. Incorporating SV data greatly improved their ability to identify strong genotype-phenotype associations, leading to the discovery of several SVs linked to sweetness and pathogen resistance.
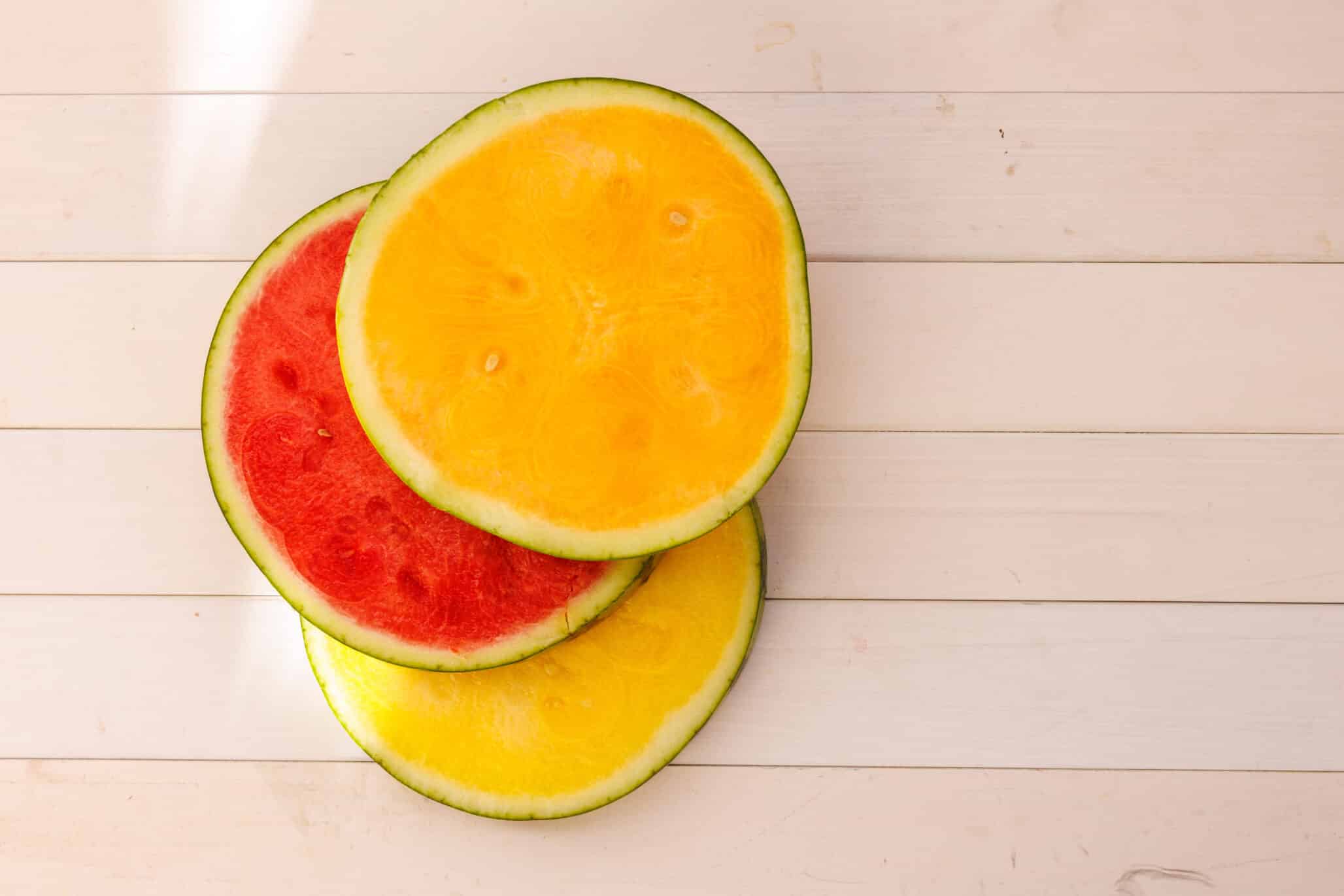
Notably, one SV was significantly associated with flesh color intensity, an important commercial trait that is linked to nutritional value and consumer appeal. This SV affected the expression of a particular gene, and as Fei stressed, “This gene could only have been identified using this super-pangenome because the SV was not linked to any nearby SNPs.”
Beyond individual gene discoveries, this super-pangenome creates opportunities for broader applications. “These resources can directly impact breeding strategies,” says Fei, “as we can use them to build platforms for genomics-assisted breeding.” To demonstrate this potential, the team developed predictive models that can infer phenotypic traits from a compact set of high-impact SNP and SV markers.
Genomic resources such as super-pangenomes offer valuable insight into the genetic basis of important traits, but their significance extends beyond the laboratory. By understanding how traits are inherited at the molecular level, researchers can design targeted breeding strategies to more efficiently develop crops with desirable combinations of traits. For watermelon, one key goal is to reintroduce disease resistance from wild relatives while preserving the sensory qualities that have made it a consumer favourite. In the era of super-pangenomes, this level of customisation is becoming increasingly achievable.