Scientists find key players in complex plant response to stress, offering clues to coping with drier conditions.
We can tell when plants need water: their leaves droop and they start to look dry. But what’s happening on a molecular level?
Scientists at the Salk Institute have made a leap forward in answering that question, which could be critical to helping agriculture adapt to drought and other climate-related stressors.
The new research suggests that in the face of environmental hardship, plants employ a small group of proteins that act as conductors to manage their complex responses to stress. The results, which are detailed in the Nov. 3 issue of Science, may help in developing new technologies to optimize water use in plants.
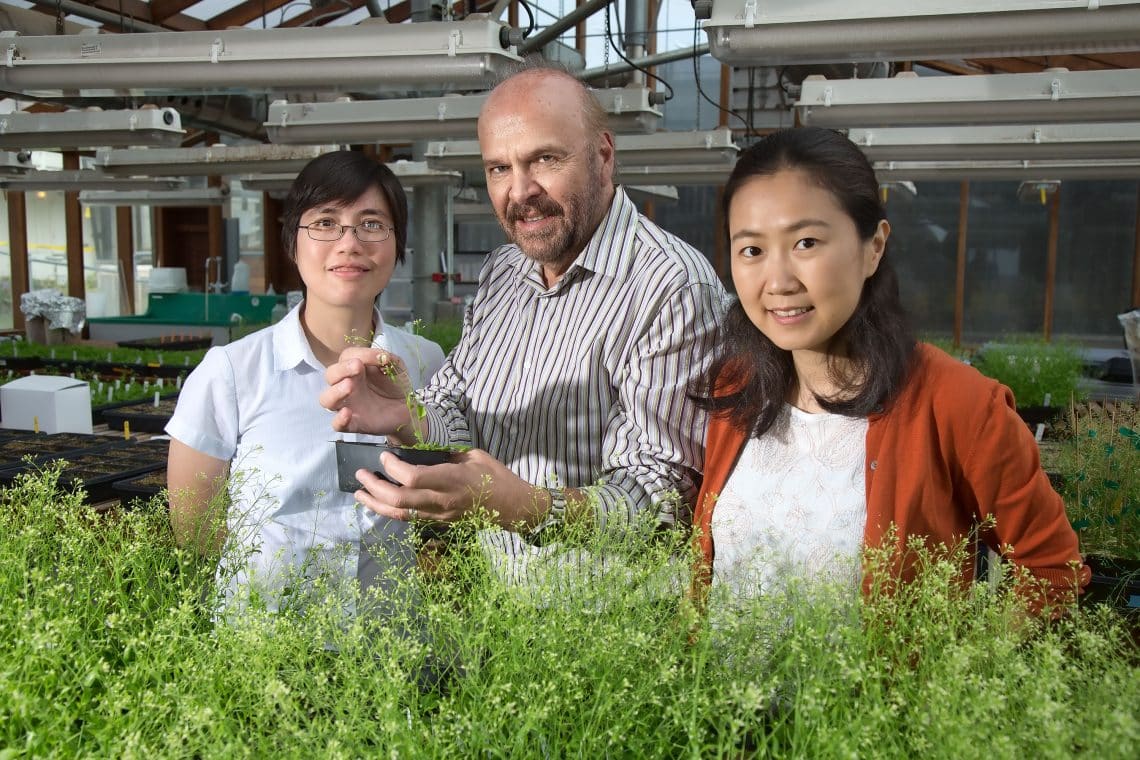
“A plant’s response to a stressor is a highly complex process at the molecular level, with hundreds of genes involved,” says senior author Joseph Ecker, a Howard Hughes Medical Institute Investigator, professor and director of Salk’s Genomic Analysis Laboratory and holder of the Salk International Council Chair in Genetics. “We’ve discovered key conductors in this molecular symphony, which may offer clues to helping plants better tolerate stressors such as drought in the face of climate change. If you can control one of these conductors, you control all of the genes that follow its lead.”
How well a plant responds to stress can determine whether it survives and thrives or succumbs to a threat. Just as humans have hormones such as adrenaline that help us cope with threats, plants have a few key hormones that allow them to respond to stressors in their environment. One of these is abscisic acid (ABA), a plant hormone involved in seed development and water optimization.
When water is scarce or salinity is high, roots and leaves produce ABA. Although the hormone is understood to impact a plant’s stress response, scientists have known very little about what happens globally after it is released.
“Just a few dozen regulatory proteins dictate the expression of hundreds if not thousands of genes,” says Liang Song, a research associate in Salk’s Plant Biology Laboratory and the paper’s first author. “By understanding what those master regulators are and how they work, we can better understand, and potentially modulate, the stress response.”
In their study, the Salk team tracked real-time changes in plant genetic activity in response to ABA and identified a handful of these master proteins that govern responses to a wide range of external stressors, including drought. Using a technique that maps where these regulatory proteins bind to DNA, the team defined key factors that coordinate gene expression, allowing for an efficient cellular response to changing conditions.
The Salk team focused on candidate regulatory proteins known to respond to ABA. They exposed 3-day-old seedlings of the reference plant Arabidopsis thaliana to abscisic acid and checked gene expression at regular time points over 60 hours.
In the process, they amassed 122 datasets involving 33,602 genes, 3,061 of which were expressed at differing levels for at least one time point. Analysis of the data revealed a hierarchy of control, with some regulatory proteins ranking as top contributors to gene expression. Intriguingly, a snapshot of protein binding patterns at a particular time point can largely explain gene expression over a large span of time. Together, these dynamics suggest a coordinated genome-wide response to environmental triggers.
“With this network view, we can see that some of these components are targeted by the same master regulator proteins, which suggests precise and coordinated genetic control,” says Song. “This could be important for agricultural purposes because regulating one gene could in turn stimulate or suppress another whole set of genes, allowing for a comprehensive design of interventions.”
The results mirror those of a 2013 study by the Ecker lab on the plant hormone ethylene, suggesting that such coordinated and hierarchical control of genetic activity may be common to flowering plants.
Other authors on the paper include: Shao-shan Carol Huang, Rosa Castanon, Joseph R. Nery, Huaming Chen, Marina Watanabe and Jerushah Thomas of the Salk Institute; and Aaron Wise and Ziv Bar-Joseph of Carnegie Mellon University.
The work was funded by the Howard Hughes Medical Institute, the Gordon and Betty Moore Foundation, the National Science Foundation, the National Institutes of Health and by a Salk Pioneer Postdoctoral Fellowship.